Cancer Proteomics and Biomarker Exploration
Cancer is a disorder resulting from autonomous, uncontrolled cell growth and differentiation, and with malignant behavior, is capable of invasion and metastasis. Carcinogenesis is initiated by non-lethal genetic damage, followed by a multi-step process involving both phenotypic and genetic changes. Regulatory genes such as the proto- oncogenes, the tumor suppressor genes, and genes regulating apoptosis are important targets of genetic damage, as well as the DNA repair genes. Mutational damage to these genes will result in activation or inactivation of the functions of their gene products, resulting in uncontrolled proliferation with abnormal differentiation and acquisition of the capability for invasion or metastasis.
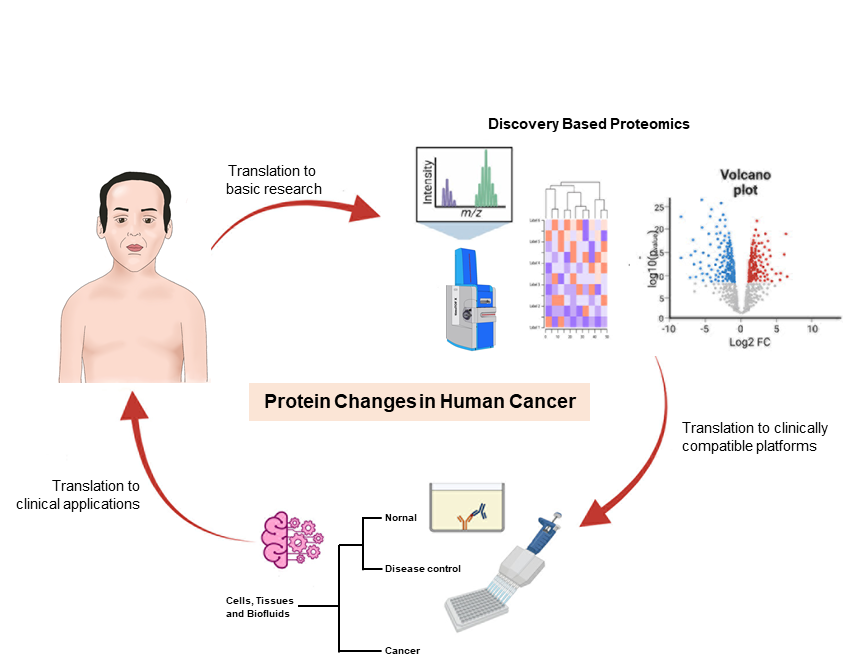
Because of our longstanding interests in proteins, we are exploring the protein changes that occur in human cancer. Initially, we collaborated with the Department of Pathology, Phramongkutklao Hospital, Bangkok to analyze changes in protein in human cancer tissues, using surgical specimens of tumor tissue and normal tissue, characterized in terms of pathology. Our group was the first research group in Thailand to use the proteomic approach in 1997, at that time using two-dimensional gel electrophoresis to compare the proteome patterns (or the total protein present at any tissue at any given time) between normal and cancer tissue. Thus, study of thyroid disease demonstrated increased expression of certain proteins, such as cathepsin B and prohibitin, in neoplastic thyroid diseases compared to non- neoplastic diseases, such as goiter or nodular hypoplasia. Immunoblotting studies also showed that metastatic potential seemed to be correlated with high expression of galectin-3.
Later as mass spectrometric studies developed, proteins of interest are positively identified by peptide mass fingerprinting, ESI/MS/MS or LC/MS-MS. We now use precision proteomics to study various systems, including protein alterations in cytosolic and secreted proteins in monolayer cell cultures, as well as three-dimensional cell culture systems which provide improved identification of secreted proteins and exosomal proteins. The mechanism(s) of cancer metastasis and cancer biomarker(s) are also being studied using precision proteomics. Thus, proteins MMRN1 and LRG1 were found for the first time, to be highly expressed urinary proteins in cervical cancer, and individual proteins LRG1 and SERPINB3 could be used to detect the occurrence of cervical cancer with 100% sensitivity and 87.5% specificity. Recently, we also employed a translational proteomic approach for the discovery, validation, and development of a multiplex cholangiocarcinoma (CCA) biomarker assay, using nine machine learning models trained on seven candidate plasma proteins. This study supports the use of the composite plasma biomarkers measured by clinically compatible ELISAs coupled to machine learning models to distinguish individuals at risk of CCA. Moreover, we identified differentially expressed proteins in the serum of advanced NSCLC patients, which respond to carboplatin plus Platinum/paclitaxel chemotherapy compared to non-responders. This finding may guide selection of treatment, so we are developing cancer biomarker assay kits for early detection and prediction of chemotherapeutic responsiveness.